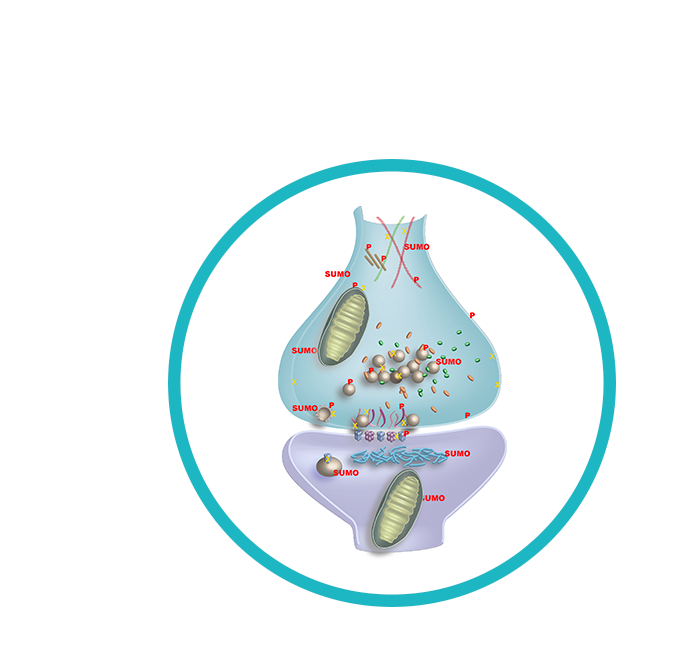
Synapses are the central information processors in the brain. Their function, efficacy and plasticity are key determinants of all brain functions, and of the corresponding behavioral output. Conversely, aberrant synapse function is the cause of many neurological and psychiatric disorders. Our ultimate objective is to generate a functional virtual synapse, in silico, that covers both the presynaptic and postsynaptic compartments.
Over the first funding period we proposed to obtain a wide set of molecular, structural and functional data on a prototypic, averaged model synapse, while only engaging a few computational projects. For the second funding period we envisioned refining these data by further wet-lab experimental work, while at the same time enhancing strongly the computational aspects, by recruiting new computational neuroscience projects into the CRC. Finally, the work will culminate in the third funding period, in which we will rely even more heavily on in silico modelling.
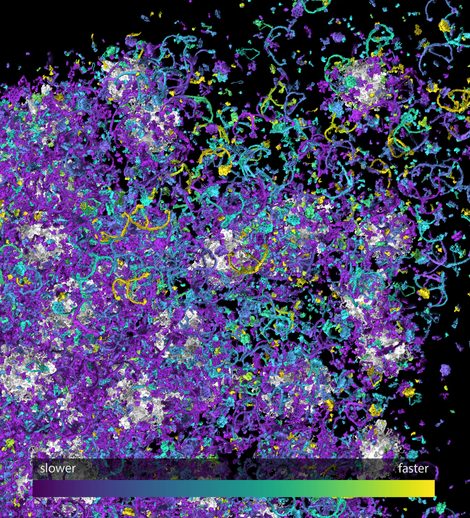
We followed this plan closely during the first funding period, and we are “on schedule” with regard to most aspects. We determined the locations of many synaptic organelles and proteins, their copy numbers, their movement rates, their metabolic turnover, several of their post-translational modifications, and their interactomes. We also performed multiple analyses of neuronal and synaptic RNAs and lipids. Finally, we initialized several modeling projects, in which collaborations with experimentalists were employed to provide an understanding of defined synaptic processes.
During the second funding period, we will continue our work along these lines, while strongly enhancing the computational aspects. As we proposed in 2017, we more than doubled the number of computational projects, which now address multiple aspects of synaptic transmission, from protein movement and nanoscale organization to long-term dynamics and plasticity. The completion of these projects will place us in an optimal position to establish synapse function models in the third funding period.
Our synaptic models will be used to answer questions on synaptic function and dysfunction, and will be extremely important in the context of connectomics efforts, since they will suggest the functional responses of synapses in given connectomes, thereby greatly increasing the functionality of structural connectomes.